 |
Quantifying Global Tolerance of Biochemical Systems: Design Implications for Moiety-Transfer Cycles
Robustness of biological organisms is widely observed although difficult to precisely characterize. Important aspects of performance can remain nearly constant within some neighborhood of the normal operating regime, leading to homeostasis, but then abruptly break down with pathological consequences beyond this neighborhood. Currently, there is no generic approach to identifying the boundaries where the local performance deteriorates abruptly and this has hampered understanding of the molecular basis of biological robustness. We, in collaboration with Dr. Armindo Salvador (University of Coimbra – Portugal) have introduced a generic approach to the characterization of boundaries between operational regimes based on the piecewise power-law representation of the system’s components. This conceptual framework allows us to define “global tolerance” as the ratio between the normal value of a parameter and the value at such a boundary. We have illustrated the utility of this concept in the context of the moiety-transfer cycle, which is a widespread module in biology. Our results show that the region of “best” local performance is surrounded by “poor” regions, and that selection for improved local performance often pushes the operating values away from regime boundaries, thus increasing global tolerance. These predictions are in agreement with experimental data from the NADPH redox cycle of human erythrocytes.
|
Starvation, the Stringent Response, and the Genome
Starvation and the stringent response of Escherichia coli and other organisms have been studied for over four decades. One known cellular response to starvation is the induction of amino acid biosynthetic pathways. However, what happens when the biosynthetic machinery under construction requires the very amino acid for which the cell is starving? To study the possible catch-22, we create a general model of cellular amino acid starvation, and we perform computational simulation and mathematical analysis to show that one possible outcome is the full failure of the system. Furthermore, we present bioinformatic data that support the evidence for cognate bias, or fewer cognate amino acids in the biosynthetic pathway, as a mechanism to avoid the quandary. Based on the initial results, we propose future work that would improve the model, define appropriate measures of bias, and use bias to draw a connection between genomic data and patterns of environmental stress. |
|
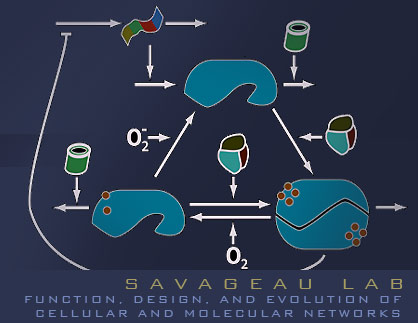
Global Regulation of E. coli Oxygen Sensing
In E. coli, FNR (for fumarate nitrate reduction) sits at the master level of respiratory regulation. Under anoxic conditions, FNR activates hundreds of genes adapting E. coli for O2 limiting growth. In an oxygen rich setting, FNR is inactive and the default state is to grow aerobically. In the aerobic state, FNR is produced and oxygen drives cycling of the protein between its three forms. The balance of the cycle favors the inactive form and serves no apparent use. Systems with this topology and apparent lack of physiological purpose have been termed futile cycles. The name itself illustrates perfectly our confusion with such systems. Our work takes a modeling approach to address the use of cycling in gene circuits, and we begin our analysis with a simple system, the FNR network.
|
Modeling Methods and Tools
As so often happens in the course of modeling, new tools and methods evolve from the work. We are identifying new methods in system analysis and optimization that aid our work. When possible, we seek to streamline and standardize the methods for use by the larger modeling community.
|
Design Space and the Design Space Toolbox
The design space of biochemical systems is a representation of the relationships between system parameters, environmental variables, and phenotypic behavior. In design space, the qualitatively distinct phenotypes of a biochemical system can be identified, counted, analyzed, and compared. Boundaries in design space indicate a transition between phenotypic behaviors and can be used to measure a system’s tolerance to large changes in parameters. Moreover, the relative size and arrangement of such phenotypic regions can suggest or confirm global properties of the system. The Design Space Toolbox provides tools to automatically construct and analyze design spaces using MATLAB.
The most recent release of the Design Space Toolbox can be downloaded here. |

